
Every environment around you – from the ground outside to the screen you’re staring at – is covered with life. Bacteria and other microbes have been found just about everywhere on Earth, often numbering in the hundreds to thousands of different species. And in fact, the health and resilience of an ecosystem, be it a rainforest or a coral reef, is closely tied to the diversity of its species. Any given species relies on its differently-skilled neighbors to provide nutrients and energy that it cannot produce by itself. Even extreme environments like the boiling hot vents at the bottom of the ocean are teeming with different microbial species.
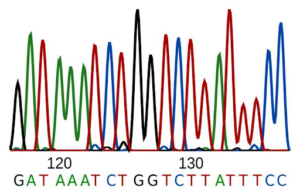
It was striking, then, in 2008 when researchers found an ecosystem containing only a single species. Published in the journal Science, the research studied water retrieved from rock fractures 2.8 km underground in the Mponeng gold mine in South Africa. After isolating DNA from the fracture water, they sequenced the DNA, meaning they determined the order of A’s, T’s, C’s, and G’s that made up the genetic code of the organisms in the water. They could then compare the sequence data to sequences previously stored in online databases to determine if the DNA came from known organisms or unknown relatives. It’s important to note that our technologies typically cannot sequence an entire piece of DNA at once – we have to sequence it in many short chunks, then use computational techniques to connect the short sequences into one or more larger DNA sequences.
When the researchers sequenced the DNA from their sample, they found that more than 99.9% of their sequences belonged to the same organism, an unnamed bacterium. For perspective, the previous record for least diverse ecosystem showed 88% of sequences belonging to one species (this was also from the Mponeng mine). In that case, you could argue that the remaining 12% of microbes could be important to the health of the ecosystem. But in the 2008 study, the remaining <0.1% was likely little more than contamination from other places in the mine and the lab.
Using the sequence data to predict what the bacterium was capable of, the researchers found that it had all the genes necessary to get the nutrition it needed from its surroundings – without relying on any other organisms. It could “breathe” sulfate and “eat” the carbon from dissolved gases like carbon dioxide and carbon monoxide, with some help from the hydrogen produced by the radioactive decay of uranium.
The researchers predicted that many of these genes were given to the bacterium horizontally by unrelated microbes in ancient times. We can imagine, then, an ancient microbe, newly equipped with skills from its current neighbors, slowly traveling deep into the underground to stake out a solitary existence. In this vein, the researchers named the species Candidatus Desulforudis audaxviator. “Audax viator” – Latin for “bold traveler” – is a reference to Jules Verne’s novel Journey to the Center of the Earth. This bold traveler shows us yet again how microbes break the rules we set for them!
Peer edited by Catalina Cobos-Uribe
