One of the most overwhelming aspects of modern-day biomedical research is the overarching heterogeneity that consumes all realms of biology. Ranging from cell to cell to human to human, we have become increasingly aware of the important differences that drive divergent responses to therapeutics and biological stimuli. The complexity of cancer is one such example.
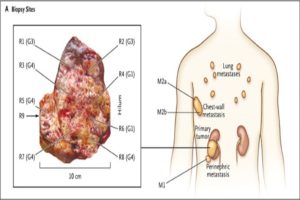
A landmark paper published by Gerlinger et al. in The New England Journal of Medicine demonstrated that analyzing multiple biopsies from a single patient’s tumor gives a much different picture of what the biology driving that tumor is, compared to examining a single biopsy alone. In addition, many studies characterizing the cellular heterogeneity of cancer have revealed that a tumor is much more than a mass of identical cells all growing out of control. Rather, a tumor is comprised of cancer cells at different stages of the cell cycle, engaged in different cell signaling pathways, as well as other cell types including immune cells and endothelial cells (the cells lining blood and lymphatic vessels).
New technologies are emerging to enable us to better dissect the heterogeneity of disease and biology. One such technique is single cell sequencing. Sequencing is a technique with which we are able to get a readout of the entire genetic composition of a cell. In its most basic form, sequencing can be performed to get a readout of DNA or RNA, which are two types of molecules involved in different levels of genetic regulation. Sequencing has traditionally been performed on bulk samples comprised of hundreds to millions of cells in aggregate. While we have made remarkable advances in our understanding of biology using bulk sequencing, the emergence of single cell sequencing has now allowed us to begin to probe the genetic profile of a single cell at varying levels of complexity. The technology often takes advantage of molecular indexing, which is a technique whereby individual mRNA or DNA molecules are labeled with a molecular tag that is associated with a single cell, and then the cells are all pooled together into one tube and sequenced. During the post-sequencing analysis, the molecular tags are re-associated with each single cell, and then the profiles of each of the single cells are compared to one another.
This novel technique has allowed us to begin investigating and understanding biology with a higher degree of resolution than we ever could have imagined before, and will undoubtedly lead to the discovery of many new exciting realms of biological regulation. For example, the biomedical company Becton and Dickson have performed a research study analyzing single cells from tumors of mouse models of cancer. They found that within each tumor sample, there were distinct populations of cells with unique gene expression profiles, and these profiles were associated with vastly different biological functions. Understanding how these different populations work together as a community to promote tumor growth may help us better understand how to develop new treatments for cancer.
However, like all cutting-edge technologies, there are still limitations that need to be overcome. One concern is inefficient collection of sample, because the amount of genetic material in a single cell is much smaller than the amount of material from a large group of cells. An additional confounding variable is uncertainty of whether a low yield of genetic information from a single cell is the result of technical error, inefficiency of small sample collection, or simply the lack of expression of a gene in that particular cell. Discerning the difference between a true, biological negative result and simply a technical deficiency are often difficult to parse out.
Especially in fields such as cancer biology, we have increasingly begun to realize that heterogeneity has largely been an obstacle in our ability to develop effective therapies and to truly understand the mechanisms of regulation of biological processes. Advances in single cell sequencing research have allowed us to further realize that there is not one function or process driving disease progression, but rather a network of cells with distinct roles. Uncoupling the forces dictating the progression of this heterogeneity is what will help us to make the next great advances in therapeutic development in cancer and many other diseases.
Peer edited by Chiungwei Huang and Richard Hodge.
Follow us on social media and never miss an article: