Most likely, you’ve taken an antibiotic before – whether it was a tablet like penicillin or a topical cream like NEOSPORIN. Antibiotics are prescribed for bacterial and fungal infections like urinary tract infections (UTIs) and athlete’s foot. Typically, antibiotics are prescribed after a bacterial or fungal infection is confirmed by your doctor, but they can also be prescribed to prevent infection, as is the case with NEOSPORIN and open wounds. But where do these antibiotics come from (besides the pharmacy)?
Around two-thirds of the antibiotics currently prescribed were actually discovered in bacteria or fungi, and the rest are man-made (synthetic). Bacteria and fungi produce antibiotics in order to compete against each other for survival. This microbial arms race started long before Sir Alexander Fleming discovered penicillin in 1928 and humans entered the fray by using bacteria’s own weapons against them. Many antibiotics are produced by bacterial and fungal enzymes called non-ribosomal peptide synthetases.
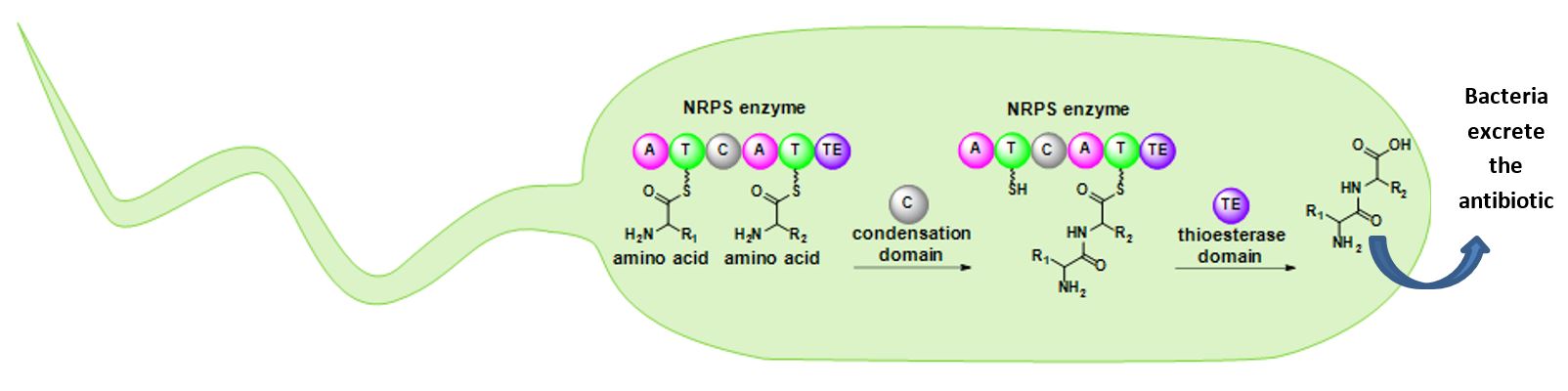
Nonribosomal peptide synthetases (NRPSs) are large enzymes that work like an assembly line to build antibiotics, pigments, toxins, and other molecules. These enzymes use amino acids as building blocks and connect them together like Lego pieces to construct molecules. NRPSs have different parts, called domains, that each have a specific job. Imagine the inside of a bacterial cell as a bucket of Legos. The adenylation (A) domain chooses a specific Lego brick (amino acid) based on size and color and passes it to the thiolation (T) domain, which holds onto the piece so that it doesn’t get lost. An NRPS typically has multiple thiolation domains. Once each thiolation domain is holding onto a Lego brick, the condensation domains click the bricks together. NRPSs can also have optional domains that further ‘decorate’ the antibiotic by performing specialized chemistry. When the antibiotic is finished, the thioesterase domain releases it from the enzyme. These incredible enzymes allow bacteria to perform complex chemistry faster and more efficiently than human chemists.
Unfortunately, bacteria have a lot of experience fighting antibiotics and quickly develop resistance. As the 2019 CDC report suggests, we are already at the crisis point and in desperate need of the next generation of antibiotics. Studying bacterial enzymes like NRPSs is one response to this call. If we understand the chemistry that bacteria use to build antibiotics, then we can engineer these enzymes to produce newer, more effective antibiotics.
Peer edited by Caitlyn Molloy
i have been doing research on weight lose from taking amoxicillin. i literally lost 15 pounds on a 10 regime. that tells me that our diet and metabolism are not the main reason for weight gain. i never changed my eating habits. as soon as i stopped taking the anti biotic my weight went back up. started again on the amoxicillin and lost another 10 pounds. what enzyme or product in the anti biotic caused the weight lose? it was the easiest way i ever lost weight.
I would like to know why?